This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
Protein Domains and Motifs
The term "protein domain" refers to the structural subunit of a protein that corresponds to a certain function or biological process [1]. Domains can be used to predict the 3-D structure of a protein and thus is its role in the organism. Domains can also be used to infer relationships between proteins. A domain is the unit used to describe a protein's motif [2]. While a motif is simply an amino acid sequence within a protein, we can use domains to envision how the motif is manifested structurally and biologically in an organism.
PER2 Domains
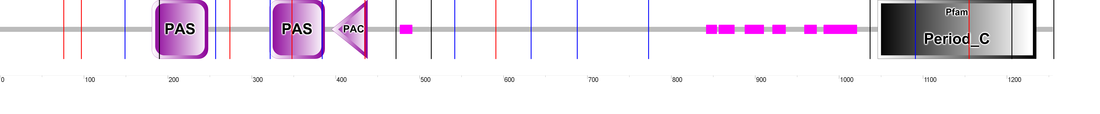
Domains as determined by SMART. The PER2 protein contains two PAS domains and a PAC motif. SMART also integrated results from Pfam, indicating PER2's protein family, Period_C. PER2 also has several regions of low complexity, as represented by the smaller, unlabeled magenta rectangles.
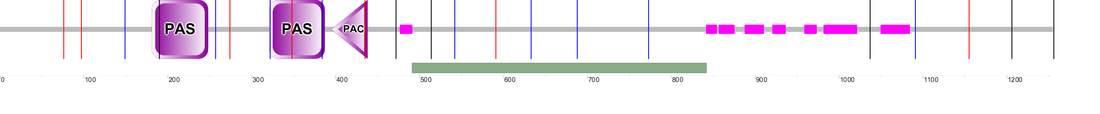
Domains as determined by Pfam. Pfam recognizes the PAS_11 domain and PER2's protein family, Period_C.
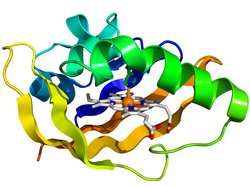
PAS:
The PAS domain (at left) consists of approximately 300 amino acids and is composed of two or more imperfect repeats. This region is involved with dimerization to a protein in the same family. This domain has been associated with light reception and regulation and circadian rhythms. In prokaryotes, the PAS domain is commonly involved with the regulations of a histidine kinase. The PAS domain is often associated with a PAC motif [3]. PAS stands for P=period circadian rhythm, A=aryl hydrocarbon receptor nuclear translocator protein, S=single minded protein [6].
The PAS domain (at left) consists of approximately 300 amino acids and is composed of two or more imperfect repeats. This region is involved with dimerization to a protein in the same family. This domain has been associated with light reception and regulation and circadian rhythms. In prokaryotes, the PAS domain is commonly involved with the regulations of a histidine kinase. The PAS domain is often associated with a PAC motif [3]. PAS stands for P=period circadian rhythm, A=aryl hydrocarbon receptor nuclear translocator protein, S=single minded protein [6].
PAC motif:
The PAC motif is always found C-terminal to PAS domains. Scientists propose that this motif aids in PAS domain folding [4].
Period_C:
This eukaryotic domain is usually between 164 and 200 amino acids long. It is associated with the PAS domain [5].
The PAC motif is always found C-terminal to PAS domains. Scientists propose that this motif aids in PAS domain folding [4].
Period_C:
This eukaryotic domain is usually between 164 and 200 amino acids long. It is associated with the PAS domain [5].
PER2 Isoforms
A protein isoform is a different form of a protein that arises from alternative splicing [7]. The isoform is produced from the same gene but undergoes a different pattern of splicing after transcription. The PER2 protein has two isoforms [8].
Isoform 1: O15055-1
- FASTA
- length= 1,255 amino acids
- mass= 136,579 da
SMART identified three of the four domains or motifs associated with the PER2 protein in isoform 1. However, the Period_C family domain is missing.
Isoform 2: O15055-2
- FASTA
- length= 404 amino acids
- mass= 45,175 da
Isoform 2 only contains one PAS domain, as identified by SMART.
Homologs
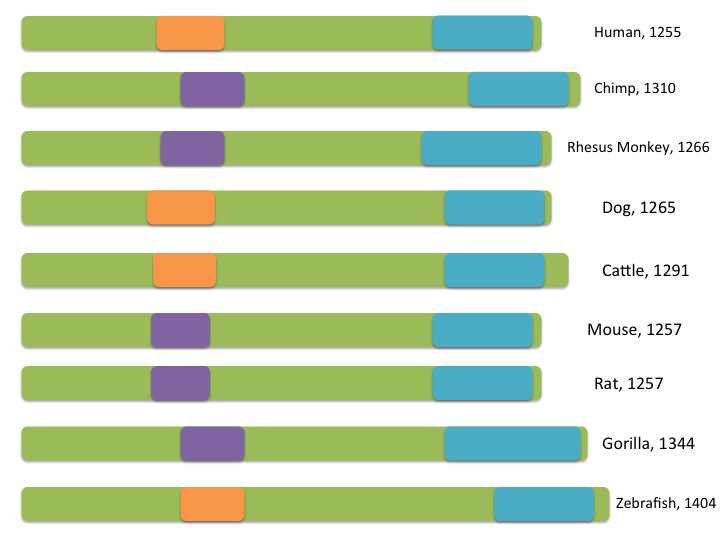
As you can see from the image above (generated with reference to HomoloGene) , the domains of the PER2 protein are highly conserved. Conservation of domains across species indicates a domain of biological significance. PAS_3 is a fold domain while the PAS domain has been found to be involved in potassium channels and binding ligands [9]. Here, we again see that the Period_C domain is associated with PAS domains. It is interesting to note that humans share their PAS domain (PAS_3) with dogs, cattle, and zebrafish, rather than primates such as chimps or gorillas.
Analysis
The online databases used for this analysis exhibited somewhat different protein structures for PER2. SMART identified two PAS domains, a PAC domain, and a Period_C domain, while Pfam and Homologene only recognized one PAS domain and the Period_C domain. Despite these differences, this information is still necessary as it implies that the PAS and Period_C domains are important for the function of PER2. PAS is responsible for light reception and subsequent regulation of the circadian rhythm. PAC may aid in folding the protein, while not much is known about the Period_C domain except that it is always found associated with a PAS domain.
The homologous proteins of PER2 are highly conserved. Conservation of domains across species indicates a domain of biological significance. While the model organisms represented here carry some variation in PAS domains, all proteins contain one PAS domain and one Period_C domain. It seems that both domains are required for normal function. The variation in PAS domains across species may indicate differences in evolution and ancestry.
The homologous proteins of PER2 are highly conserved. Conservation of domains across species indicates a domain of biological significance. While the model organisms represented here carry some variation in PAS domains, all proteins contain one PAS domain and one Period_C domain. It seems that both domains are required for normal function. The variation in PAS domains across species may indicate differences in evolution and ancestry.
References
[1] EMBL-EBI Train Online. What are Protein Domains?
[2] Richardson, J. (1994). Introdutction: Protein Motifs. The FASEB Journal, 8(15).
[3] PROSITE. (2001). PAS repeat and PAC domain profiles. Retrieved February 24, 2014, from http://prosite.expasy.org/cgi-bin/prosite/nicedoc.pl?PS50112
[4] SMART: http://smart.embl-heidelberg.de/smart/show_motifs.pl
[5] Pfam: http://pfam.sanger.ac.uk/family/PF12114.3
[6] Pfam: http://pfam.sanger.ac.uk/family/PF08447
[7] MeSH. Protein isoforms. Retrieved February 24, 2014, from http://www.ncbi.nlm.nih.gov/mesh?cmd=Retrieve&dopt=Full&list_uids=68020033
[8] GeneCards
[9] HomoloGene: http://www.ncbi.nlm.nih.gov/homologene/?term=NP_073728
[1] EMBL-EBI Train Online. What are Protein Domains?
[2] Richardson, J. (1994). Introdutction: Protein Motifs. The FASEB Journal, 8(15).
[3] PROSITE. (2001). PAS repeat and PAC domain profiles. Retrieved February 24, 2014, from http://prosite.expasy.org/cgi-bin/prosite/nicedoc.pl?PS50112
[4] SMART: http://smart.embl-heidelberg.de/smart/show_motifs.pl
[5] Pfam: http://pfam.sanger.ac.uk/family/PF12114.3
[6] Pfam: http://pfam.sanger.ac.uk/family/PF08447
[7] MeSH. Protein isoforms. Retrieved February 24, 2014, from http://www.ncbi.nlm.nih.gov/mesh?cmd=Retrieve&dopt=Full&list_uids=68020033
[8] GeneCards
[9] HomoloGene: http://www.ncbi.nlm.nih.gov/homologene/?term=NP_073728