This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
Homology
In 1848, Richard Owen coined the term "homology" to define the physical similarities of organisms [1]. Homologous structures indicate a common evolutionary ancestor. An example of homology is the forelimb of tetrapods. Frogs, rabbits, lizards, and birds have forelimbs with different functions, but these forelimbs all contain the same bones--bones that were unearthed in a fossil of an extinct common ancestor [2]. Like physical features such as appendages, DNA homology can also be used to determine the probability of two genes sharing a common ancestor [3]. However, there is one caveat. One must be wary of analogs. Analogous sequences or physical similarities can result from convergent evolution or chance [3].
The homologs of PER2 were determined using the National Center for Biotechnology Information (NCBI)'s program, BLAST. Basic Local Alignment Search tool, or BLAST allows one to enter the FASTA sequence of DNA, mRNA, or protein. BLAST uses a database to compare these sequences, identifying similarities and identical sequences in other organisms. The larger the percent of similarity, the lower the probability that these sequences are similar due to convergent evolution or chance [4]. Since the PER2 gene is large, I chose to input its mRNA sequence into BLAST to search for homologs. BLAST indicated quite a few homologs (see below). These results were confirmed using Homolgene. Please note that the list of homologs (below) is not extensive, but rather one containing the homologs with the most similar sequences and homologs in model organisms often used in genetic research.
The homologs of PER2 were determined using the National Center for Biotechnology Information (NCBI)'s program, BLAST. Basic Local Alignment Search tool, or BLAST allows one to enter the FASTA sequence of DNA, mRNA, or protein. BLAST uses a database to compare these sequences, identifying similarities and identical sequences in other organisms. The larger the percent of similarity, the lower the probability that these sequences are similar due to convergent evolution or chance [4]. Since the PER2 gene is large, I chose to input its mRNA sequence into BLAST to search for homologs. BLAST indicated quite a few homologs (see below). These results were confirmed using Homolgene. Please note that the list of homologs (below) is not extensive, but rather one containing the homologs with the most similar sequences and homologs in model organisms often used in genetic research.
PER2 (Period Circadian Clock 2) Gene Homologs
Human (Homo sapiens): PER2, period circadian clock 2
Accession number: NC_000002.12 GI Number: 568815596 FASTA mRNA 
Rhesus Monkey (Macaca mulatta): PER2
Accession Number: XM_001086845.2 GI Number: 297265212 FASTA E Value: 0.0 Max Identical: 96% 
Cattle (Bos taurus): PER2
Accession Number: NM_001192317.1 GI Number: 300796795 FASTA E Value: 0.0 Max Identical: 81% 
Brown Rat (Rattus norvegicus): mRNA for rPer2
Accession Number: NM_031678.1 GI Number: 13928941 FASTA E Value: 0.0 Max Identical: 79% |

Chimpanzee (Pan troglodytes): transcript variant 2 (PER2)
Accession number: XM_001154734.3 GI Number: 410036398 FASTA E Value: 0.0 Max Identical: 99% 
Dog (Canis lupis familiaris): transcript variant X3 (PER2)
Accession number: XM_005635927.1 GI Number: 545543072 FASTA E Value: 0.0 Max Identical: 81% 
House Mouse (Mus musculus): Per2
Accession number: NM_011066.3 GI Number: 153792235 FASTA E Value: 0.0 Max Identical: 80% 
Western Lowland Gorilla (Gorilla gorilla gorilla): PER2
Accession number: XM_004033434.1 GI Number: 426339058 FASTA E Value: 0.0 Max identical: 99% |
Analysis
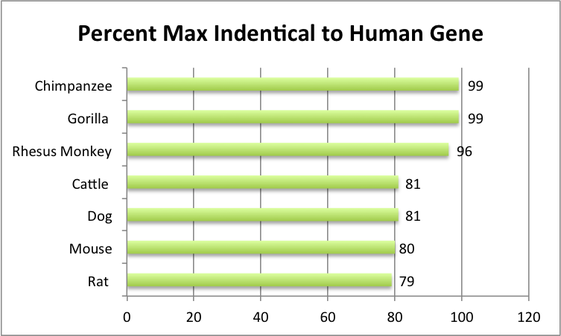
As you can see from this brief list of homologs, the PER2 gene is highly conserved among mammals. This homology reflects the importance of the PER2 gene. Since PER2 regulates the circadian rhythm, an essential cycle in living organisms, this is not surprising. The PER2 genes of the three primates represented here are almost identical to the human PER2 gene. This could suggest a common ancestor. Explore the phylogeny page for more information on these homologs relationships.
References
[1] Evolution Makes Sense of Homologies. http://www.zoology.ubc.ca/~bio336/bio336/lectures/lecture5/overheads.html
[2] Understanding Evolution. Homologies.
[3] Schope, Thomas. (1983). Homology. Science,219 (4582). http://www.jstor.org/stable/1690261
[4] Koonin, E.V. and Gaperin, M.Y. Sequence-Evolution-Function: Computational Approaches in Comparative Genomics. Boston: Kluwer Academic; 2003. Chapter 2, Evolutionary Concept in Genetics and Genomics. Available from http://www.ncbi.nlm.nih.gov/books/NBK20255/
[1] Evolution Makes Sense of Homologies. http://www.zoology.ubc.ca/~bio336/bio336/lectures/lecture5/overheads.html
[2] Understanding Evolution. Homologies.
[3] Schope, Thomas. (1983). Homology. Science,219 (4582). http://www.jstor.org/stable/1690261
[4] Koonin, E.V. and Gaperin, M.Y. Sequence-Evolution-Function: Computational Approaches in Comparative Genomics. Boston: Kluwer Academic; 2003. Chapter 2, Evolutionary Concept in Genetics and Genomics. Available from http://www.ncbi.nlm.nih.gov/books/NBK20255/