This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
Protein Motifs
Protein motifs consist of a sequence of amino acids. When in common, they can act as signatures of protein families and can often be used to predict protein function [2]. While a motif is simply an amino acid sequence within a protein, it may code for a protein domain. The term "protein domain" refers to the structural subunit of a protein that corresponds to a certain function or biological process [3]. Domains can be used to predict the 3-D structure of a protein and thus is its role in the organism. Domains can also be used to infer relationships between proteins. A domain is the unit used to describe a protein's motif [4].
MEME
The NCBI database, MEME, was used to discover conserved protein motifs among the PER2 homologs. According to their website, the MEME suite and its algorithms allow you to input unaligned DNA, mRNA, or protein sequences and retrieve motifs among these sequences. MEME also compares motifs to other motifs and matches motifs with Gene Ontology terms. MEME identified three motifs among the PER2 homologs. For the full MEME job, click here.
| per2_protein_homolog_fastas.docx | |
| File Size: | 167 kb |
| File Type: | docx |
How to read a Motif
|
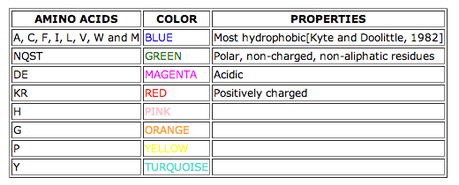
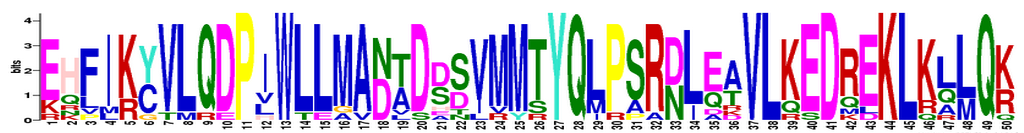
The following instructions from the MEME suite succinctly explain how to read a motif: "MEME motifs are represented by position-specific probability matrices that specify the probability of each possible letter appearing at each possible position in a occurrence of the motif. These are displayed as 'sequence LOGOS', containing stacks of letters at each position in the motif. The total height of the stack is the 'information content' of that position in the motif in bits. The height of the individual letters in a stack is the probability of the letter at that position multiplied by the total information content of the stack" [5]. Each letter represents an amino acid, which are identified in the chart at right [6].
|
Motif 1
Starting at Homo sapiens amino acid 317.
Motif 2
Starting at Homo sapiens amino acid 395.
Motif 3
Starting at Homo sapiens amino acid 1125.
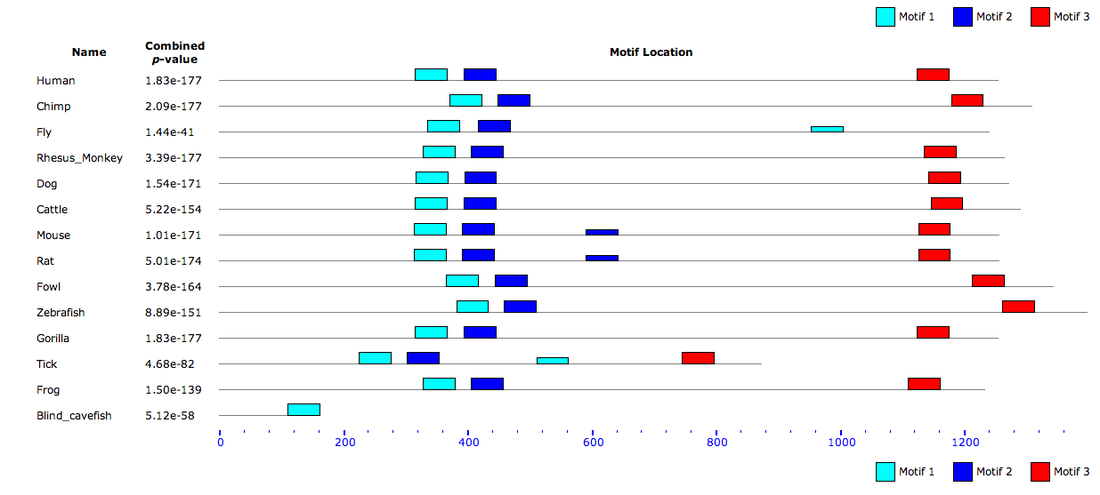
Motif Locations
Analysis
Of the three motifs, motif 1 is the most conserved among homologs. We can tell this because this motif has the largest letters and least number of letters per position. Motif 3 is the least conserved; it has the most variance in amino acids at each position. If you look at the location of each motif (above), you can see that for the most part, the location of each motif amid the homolog proteins is also fairly well conserved. This conservation may indicate that these motifs are essential for the function of the PER2 protein. This makes sense as PER2 is a circadian rhythm protein whose function is conserved among its homologs.
References
[1] Bailey, T.L., and Elkan, C. (1994). Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology. pp. 28-36, AAAI Press, Menlo Park, California.
[2] Bork, P. and Koonin, E.V. (1996). Protein Sequence Motifs. Current Opinion in Structural Biology. 6(3): 366-376. doi: 10.1016/S0959-440X(96)80057-1
[3] EMBL-EBI Train Online. What are Protein Domains?
[4] Richardson, J. (1994). Introdutction: Protein Motifs. The FASEB Journal, 8(15).
[5] MEME. http://nbcr-222.ucsd.edu/opal-jobs/appMEME_4.9.113999291784481108528430/meme.html#summary_doc
[6] Kyte, J. and Doolittle, R.J. (1982). A Simple Method for Displaying the Hydropathic Character of a Protein. J. Mol Biol. 157, 105-132.
[1] Bailey, T.L., and Elkan, C. (1994). Fitting a mixture model by expectation maximization to discover motifs in biopolymers. Proceedings of the Second International Conference on Intelligent Systems for Molecular Biology. pp. 28-36, AAAI Press, Menlo Park, California.
[2] Bork, P. and Koonin, E.V. (1996). Protein Sequence Motifs. Current Opinion in Structural Biology. 6(3): 366-376. doi: 10.1016/S0959-440X(96)80057-1
[3] EMBL-EBI Train Online. What are Protein Domains?
[4] Richardson, J. (1994). Introdutction: Protein Motifs. The FASEB Journal, 8(15).
[5] MEME. http://nbcr-222.ucsd.edu/opal-jobs/appMEME_4.9.113999291784481108528430/meme.html#summary_doc
[6] Kyte, J. and Doolittle, R.J. (1982). A Simple Method for Displaying the Hydropathic Character of a Protein. J. Mol Biol. 157, 105-132.