This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
PER2 in Humans
Overview [2]
The PER2 protein regulates itself via indirect negative regulation (it cannot directly bind to DNA). It has two transcriptional activators, Clock and Bmal1. Clock and Bmal1 form a heterodimer, Clock-Bmal1. Clock-Bmal1 binds to the PER2 promoter, initiating expression. PER2 enters the nucleus and disrupts the actions of Clock-Bmal1, though its exact mechanism is unknown.
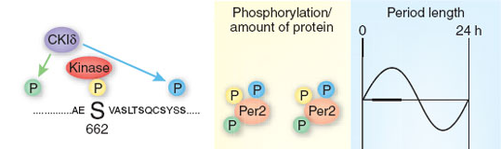
Once transcribed, the PER2 protein is phosphorylated (see image above). An unknown kinase phosphorylates the protein at a serine residue (Ser662). This initial phosphorylation allows casein kinase Iε (CKIε) to phosphorylate a neighboring residue (blue in image), as well independently phosphorylating additional residues at the same time (green). This pattern of phosphorylation produces PER2 protein at intermediate levels.
A mutation in CKIε or in the serine residue (S662G) in the PER2 gene and subsequent protein has been linked to a familial advanced sleep phase syndrome (FASPS), a syndrome of circadian nature. This mutation causes hypophosphorylation in PER2 and lower levels of the protein. In mice models it was determined that PER2 protein abundance affected the circadian rhythm. Lower levels of protein correlated with a shorter circadian cycle. This study provided insight on the regulation of the PER2 protein and may have implications for the role of PER2 in depression, which has genetics links to our circadian rhythm.
Once transcribed, the PER2 protein is phosphorylated (see image above). An unknown kinase phosphorylates the protein at a serine residue (Ser662). This initial phosphorylation allows casein kinase Iε (CKIε) to phosphorylate a neighboring residue (blue in image), as well independently phosphorylating additional residues at the same time (green). This pattern of phosphorylation produces PER2 protein at intermediate levels.
A mutation in CKIε or in the serine residue (S662G) in the PER2 gene and subsequent protein has been linked to a familial advanced sleep phase syndrome (FASPS), a syndrome of circadian nature. This mutation causes hypophosphorylation in PER2 and lower levels of the protein. In mice models it was determined that PER2 protein abundance affected the circadian rhythm. Lower levels of protein correlated with a shorter circadian cycle. This study provided insight on the regulation of the PER2 protein and may have implications for the role of PER2 in depression, which has genetics links to our circadian rhythm.
Interactions [1]
|
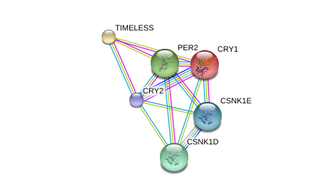
PER2 interacts directly with the other period proteins, PER1 and PER3. Via its C-terminal domain, PER2 interacts with CRY1 and CRY2. Interaction with CSNK1D and CSNK1E spurs nuclear localization of PER proteins. PER2 interacts with dozens of more proteins. An image showing all known associations can be found here.
PER2 has six posttranslational modification (PTM) sites [5]. All six are phosphorylation sites at the amino acid, serine. Five occur between the 662nd and 674th positions, while the sixth occurs at the 977th position [6]. |
Isoforms
A protein isoform is a different form of a protein that arises from alternative splicing [3]. The isoform is produced from the same gene but undergoes a different pattern of splicing after transcription. The PER2 protein has two isoforms [1].
Isoform 1: O15055-1
Isoform 2: O15055-2
A protein isoform is a different form of a protein that arises from alternative splicing [3]. The isoform is produced from the same gene but undergoes a different pattern of splicing after transcription. The PER2 protein has two isoforms [1].
Isoform 1: O15055-1
- FASTA
- length= 1,255 amino acids
- mass= 136,579 da
Isoform 2: O15055-2
- FASTA
- length= 404 amino acids
- mass= 45,175 da
References
[1] GeneCards
[2] Albrecht, Urs. (2007). Per2 has time on its side. Nature Chemical Biology. 3, 139-140. doi:10.1038/nchembio0307-139
[3] MeSH. Protein isoforms. Retrieved February 24, 2014, from http://www.ncbi.nlm.nih.gov/mesh?cmd=Retrieve&dopt=Full&list_uids=68020033
[4] NCBI: Gene. http://www.ncbi.nlm.nih.gov/gene/8864
[5] SMART http://smart.embl-heidelberg.de/smart/show_motifs.pl
[6] PTMcode
[1] GeneCards
[2] Albrecht, Urs. (2007). Per2 has time on its side. Nature Chemical Biology. 3, 139-140. doi:10.1038/nchembio0307-139
[3] MeSH. Protein isoforms. Retrieved February 24, 2014, from http://www.ncbi.nlm.nih.gov/mesh?cmd=Retrieve&dopt=Full&list_uids=68020033
[4] NCBI: Gene. http://www.ncbi.nlm.nih.gov/gene/8864
[5] SMART http://smart.embl-heidelberg.de/smart/show_motifs.pl
[6] PTMcode