This web page was produced as an assignment for Genetics 564, an undergraduate course at UW-Madison
Ontology
Gene ontology (GO) is the language used by scientists to classify genes and their products. Keywords and lists are used for description and grouping [1]. The Gene Ontology program employs three different ontologies to categorize genes [1].
GO is a dynamic program that is updated every day as more research is conducted [1]. Gene ontology is important for organizing and understanding the many components of a gene and its products.
- molecular function= biochemical activity of the gene's protein
- biological process= the overall biological purpose and function of the gene's protein
- cellular component= where in the cell the gene's protein localizes
GO is a dynamic program that is updated every day as more research is conducted [1]. Gene ontology is important for organizing and understanding the many components of a gene and its products.
Term Associations of PER2 [2]
|
Molecular Function
|
Biological Process
|
Cellular Component
|
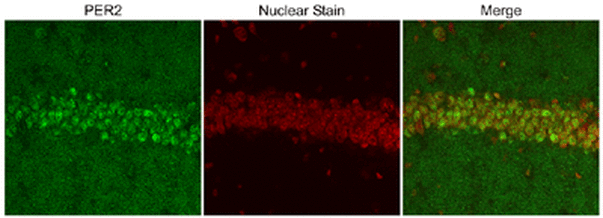
The above image is a series of pictomicrographs of PER2 localization in hippocampus cells in mice. The left panel exhibits the staining pattern of PER2 antibodies. The middle panel exhibits the staining pattern of the nuclear marker. When these images are merged (in the right panel), we can see that the staining of PER2 overlaps with the staining of the nucleus. Additionally, you can see some immunoreactivity that did not localize to the nucleus, indicating that PER2 also resides in the cytoplasm [3].
Analysis
The term associations produced by The Gene Ontology program are not surprising. PER2 is involved in circadian rhythm maintenance and regulates its own transcription. The PER2 protein regulates itself via indirect negative regulation (it cannot directly bind to DNA). It has two transcriptional activators, Clock and Bmal1. Clock and Bmal1 form a heterodimer, Clock-Bmal1. Clock-Bmal1 binds to the PER2 promoter, initiating expression. PER2 enters the nucleus and disrupts the actions of Clock-Bmal1, though its exact mechanism is unknown [4]. In this way, we would expect "nucleus" as a GO term. PER2 localized in the "cytoplasm" may be awaiting its transcriptional activators before moving into the nucleus of cells in the brain.
References
[1] Hill, D., et al. (2010). Representing Ontogeny Through Ontology: A Developmental Biologist's Guide to The Gene Ontology. Mol Reprod Dev. 77(4), 314-329. doi: 10.1002/mrd.21130
[2] AmiGO http://amigo.geneontology.org/cgi-bin/amigo/gp-assoc.cgi?gp=UniProtKB:O15055&session_id=9333amigo1392936779
[3] Wang, L., et al. (2009). Expression of the circadian clock gene Period2 in the hippocampus: possible implications for synaptic plasticity and learned behavior. ASN Neuro. Jun 10;1(3). pii: e00012. doi: 10.1042/AN20090020
[4] Albrecht, Urs. (2007). Per2 has time on its side. Nature Chemical Biology. 3, 139-140. doi:10.1038/nchembio0307-139
[1] Hill, D., et al. (2010). Representing Ontogeny Through Ontology: A Developmental Biologist's Guide to The Gene Ontology. Mol Reprod Dev. 77(4), 314-329. doi: 10.1002/mrd.21130
[2] AmiGO http://amigo.geneontology.org/cgi-bin/amigo/gp-assoc.cgi?gp=UniProtKB:O15055&session_id=9333amigo1392936779
[3] Wang, L., et al. (2009). Expression of the circadian clock gene Period2 in the hippocampus: possible implications for synaptic plasticity and learned behavior. ASN Neuro. Jun 10;1(3). pii: e00012. doi: 10.1042/AN20090020
[4] Albrecht, Urs. (2007). Per2 has time on its side. Nature Chemical Biology. 3, 139-140. doi:10.1038/nchembio0307-139